Have you ever wanted to simulate a whole cell, just on your laptop?
No lab bench, no pipetting, goodbie gloves… It would be incredible! So, are virtual cells here? Read on to find out!
Don’t keep this newsletter a secret: Forward it to a friend today!
Was this email forwarded to you? Subscribe here!
Cells Go Virtual
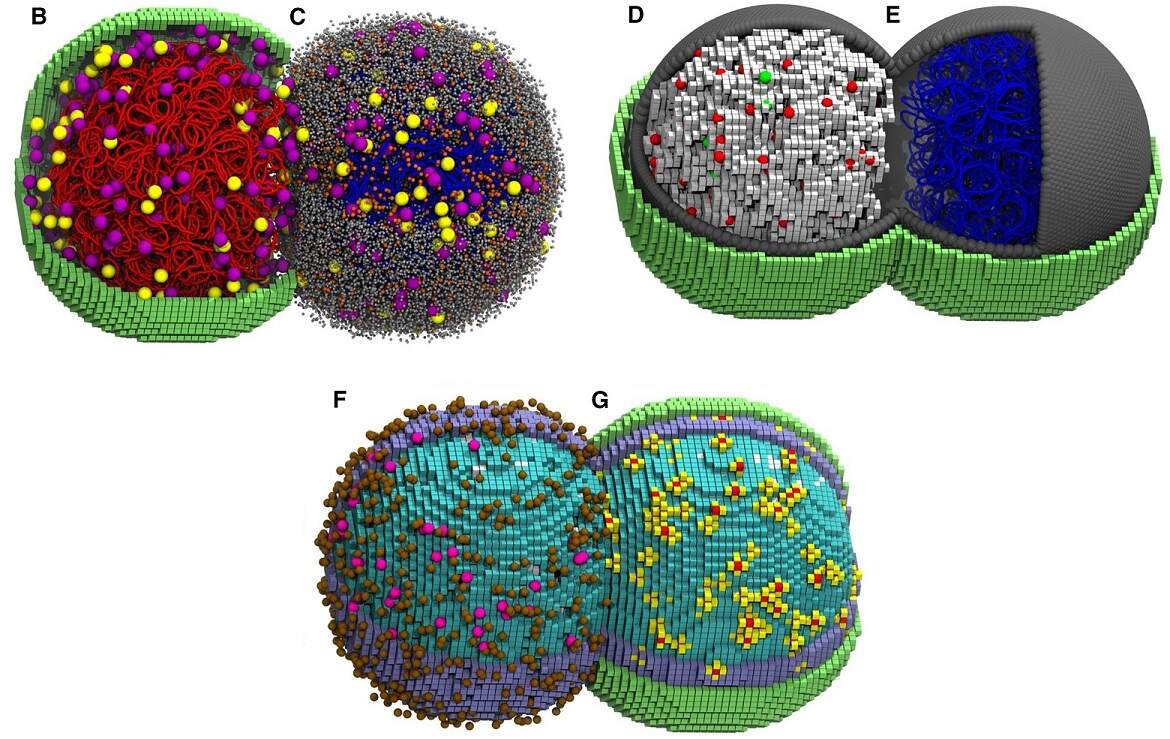
Scientists created a whole-cell simulation of a small, synthetic bacterium, accurate in time and space, recreating biological traits from the ground up. Image credits: Cell Press.
Simulating Full Cells
What does it take to simulate an entire cell on a computer?
I know, it’s one of those questions you ask at 2 a.m after one too many at the university bar. But some researchers are taking it seriously! The idea of a “virtual cell” has been around for at least 20 years. And probably even before!
The concept is simple. Build a computational model that reproduces how a cell behaves, starting from its molecular components.
Why, you ask?
Well, first of all, it’s super cool. In addition, it would be game-changing for basically every field of biology! You design your experiments, and then just run them on your computer!
No lab work → money and time saved.
Imagine a molecular biologist testing how a mutation affects cell growth in seconds. Or predict drug toxicity just on a computer! Or build powerful biomanufacturing systems…
The possibilities are endless!
Where Are Our Virtual Cells?
But how does a virtual cell actually look?
In practice, it’s equations that build a bottom-up simulation of cellular processes. You take experimental data and model all biochemical and physical processes together!
Then, you look: does it behave like a real cell?
3 things are important for a virtual cell model:
Completeness → It represents the whole cell
Dynamics → It represents cellular behaviours like size or cell cycle, over time
Predictive → It works under different conditions!
Amazing! Where are our virtual cells?
About that…
It’s hard; there is no way around it. To create a virtual cell, you need to understand it completely. We’re talking about every reaction and interaction, quantitatively, in space and time!
And by the way, we can’t even measure that experimentally. There is no way to observe a full cell all at once! We always rely on bits and pieces of data.
Of course, this didn’t stop scientists from trying.
There are 3 main ways to create a virtual cell model:
Static structural models: Think atomic simulations. They have insane resolution (down to atoms!), but they’re static and don’t capture dynamics.
Well-stirred kinetic models: They assume that cells are homogeneous solutions (which is not true!) and capture average reaction dynamics.
AI models: The new kid on the block. They have some predictive power, but tell you nothing about the biology.
So, static models are too short in time to cover gene expression or cellular division, and well-stirred models miss local diffusion and random encounters in 3D. And AI models are still too black-boxy!
Are we out of options?
Looking at Cells in 4D
Of course, there is hope! It’s in today’s paper.
The authors built a 4D whole-cell model (space + time) of the minimal bacterium JCVI-syn3A (catchy name). This model combined simulations at different resolutions to capture cell behaviour for almost 2 hours!
And the 50 virtual cells they simulated don’t just look pretty. They actually reproduce key experimental data: doubling time, DNA replication rates, and cell division!
What did they simulate? Glad you asked.
Syn3A: The Perfect Test Cell
JCVI-syn3A (or just Syn3A) is a genetically minimal, semi-synthetic bacterium, engineered to contain the smallest number of genes necessary for independent life.
Its genome is based on Micoplasma mycoides, and it has only:
493 genes
On a ~543 kbp genome
So small! And crucially, it’s well characterized:
Annotated genome
Proteomics and transcriptomic dataset
Morphology data from cryo microscopy
DNA sequencing data and chromosome contact maps
Small and well-known: perfect for a full-cell simulation!
Combining Different Resolution Models
This is a hybrid model.
No single modeling approach can handle everything in a cell. So, the team combined several computational methods to handle different biological processes.
They used:
RDME (reaction-diffusion master equation): Handles spatial reactions and diffusion in 3D. Used for translation, RNA degradation, and more!
CME (chemical master equation): Equations to represent the stochastic kinetics of transcription and tRNA dynamics.
ODEs (ordinary differential equations): Classic metabolic modeling; they are used for glycolysis, nucleotide synthesis, and lipid metabolism.
Brownian dynamics: It handles DNA and chromosome physics: replication, motion, and binding proteins!
Each cell runs for 2 hours of biological time. But real-world time? Each replicate takes 4-6 days of computation, and about 250 GPU hours! And they simulated 50 cells!
An insane amount of computation. Not cheap either!
Spatial Components: The Key Factor
The big upgrade over previous models?
This one treats the cell as a real, spatial environment. Cells are not well-stirred reactors, even if it simplifies calculations. In cells, molecules are present in low copies and organized in 3D spaces. To interact, they need to find each other!
This spatial heterogeneity affects everything:
Replication initiation
Transcription
Translation
mRNA degradation
and more!
So, what’s inside this messy environment?
Almost all cellular components are here:
DNA is represented as a low-resolution polymer
RNA and proteins are particles
Ribosomes have a special representation, because they are too large for the lattice
Membrane proteins and lipids can insert into the membrane to grow it
Chromosome position changes dynamically, and affects transcription!
Predicting Experimental Results
So, how do you check if your virtual cell is working?
Well, you compare to experimental results. And it worked!
Doubling time
The simulation predicted a membrane doubling time of ~105 min, which matched exactly the experimental doubling time! And it beat the earlier well-stirred model, which predicted 97 min.DNA replication timing
DNA replication in bacteria is divided into 3 periods (B, C, and D; don’t ask about A). The model predicts:B period ~5 min
C period ~46 min
D period ~54 min
These align with experiments!
Cell growth and division
The model links membrane growth and lipid metabolism. When compared with microscopy data, the morphology distribution was quite similar! Even if not perfect.
Are Virtual Cells Here?
Such an amazing work!
There’s so much to unpack here, I can’t do it all! It’s an exciting direction. There has been a lot of hype around virtual cells, especially with AI.
So, are we there?
Not even close.
I’m super excited, but it’s hard not to see how far any of this is from real use. It has just so many limitations:
Entire biological machines missing: the big ones are polyribosomes, resulting in a lower-than-expected protein production.
Simplified chromosome segregation: They need to add a small force, or cells don’t divide.
Limited data: Probably the biggest problem. Syn3A is a small and simple organism. Even for that, we don’t have complete data! They had to combine data from different bacteria.
Computational costs: They used an insane amount of computation to simulate the smallest organism we know of. The second biggest problem!
So, yeah, we’re not getting virtual cells anytime soon.
But this is still a great step ahead! A simulation that represents space and time, for a whole cell? And it simulates up to 2 hours?! That’s incredible!
And maybe there are some niche applications where something like this could already be helpful. Maybe to improve on existing metabolic engineering…
But go and read the paper, it’s a cool one! And judge for yourself.
If you made it this far, thank you! What do you think of virtual cells?? Do you think we’re ever going to be able to do it? Reply and let me know!
P.S: Know someone interested in SynBio? Share it with them!
What did you think of today's newsletter?
More Room:
Low-Temp DNA Origami: DNA origami is cool, but you need high temperature for synthesis, and that’s a big drawback. In this study, the authors show that adding glycerol enables DNA self-assembly at arbitrary temperatures, even below freezing, without denaturation. The method supports diverse structures and is compatible with proteins from cell lysates, offering a simple approach for DNA nanotechnology in physiological settings.
Shining Light on Ear’s Proteins: Stereocilia rely on myosins to transport components needed for hearing, but their movement is not clear. In this study, the authors develop STELLA-SPIM microscopy to track single myosin molecules in live cells and show that they move processively toward stereocilia tips when activated or dimerized. When constrained by interactions, it shifts to non-processive, step-like movement. These results suggest that release from autoinhibition enables MYO7A-driven transport in stereocilia.
Stronger DNA Origami? Another problem with DNA origami? It’s more sensitive than non-bio materials. DNA–silica composites are promising, but controlling silica thickness is hard. In this study, the authors tune DNA-templated silicification, leading to thinner, more uniform silica coatings. This approach enables more control, improves monodispersity, and allows surface functionalization, advancing DNA-based materials for nanotech applications.
