DNA nanotech can do many things.
Can it help us understand how the cytoskeleton assembles inside cells? Well, all-DNA synthetic cells might do the trick! Cytoskeleton research meets DNA nanotech.
Don’t keep this newsletter a secret: Forward it to a friend today!
Was this email forwarded to you? Subscribe here!
Science 🤝 Podcast: I’m a big podcast fan. You tell me I can learn while commuting? Amazing! So, I was excited to discover Boba & Biotech! This new podcast explored what it takes to develop new drugs and the ecosystem of academia, biotech, and investors behind it. My favourite is episode 3. Life science tools are an undervalued part of the ecosystem. Plus, an interesting discussion about the value (or lack of) animal models, from the perspective of a trained veterinarian! Go and have a listen!
DNA Builds a Cytoskeleton
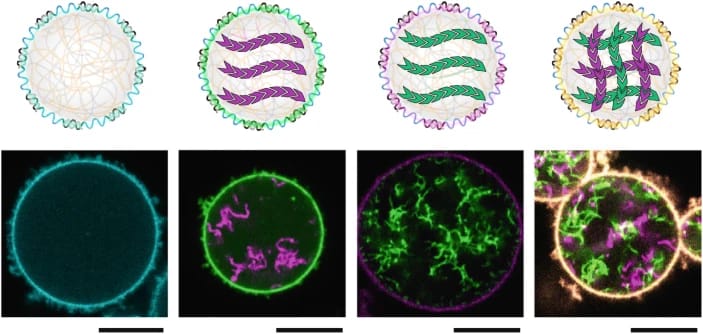
Researchers built all-DNA synthetic cells to study the assembly of artificial cytoskeletons, creating a programmable and versatile platform. Image credits: Nature.
Cells are busy places.
Inside the cytoplasm, there’s a crowded and complex universe. It’s easy to forget (at least for me), but everything inside cells happens in extremely packed environments.
At any given moment, many things are happening: nutrients are exchanged, proteins are produced, and signals are exchanged. And more! Cells pull this off by compartmentalizing different functions and structures.
Put together organelles, cytoskeletons, and macromolecules, and they occupy 40% of cellular volume! That leads to 2 key properties of the cytoplasm:
Crowdness: Closer molecules → different reaction behaviors. Like increasing concentration without adding anything.
Viscoelasticity: Cells behave like something between a fluid and a solid. Thanks to the cytoskeleton, they resist deformation and adapt to stress.
These two properties are closely linked and influence how cellular systems work. Take the cytoskeleton: how does it form in an environment that is not only crowded, but even pushing back?
And how do you even study that?!
Synthetic Cells to the Rescue
One answer: synthetic cells.
Synthetic cells (SCs) are one of the dreams of synthetic biologists. If you can build a cell, you can control everything inside it. In theory, biology would get close to engineering, transforming drug delivery, biosensing, and biomanufacturing.
But we’re far from there! So, you don’t have to worry.
There are two main ways that you can take:
Top-down: Take a cell and simplify it.
Bottom-up: Start with non-living blocks and assemble them into something cell-like.
For me, the bottom-up approach is so much cooler! Plus, bottom-up synthetic cells are perfect for testing systems that are too complex to study in real cells.
All-DNA Synthetic Cells
This brings us to today’s paper!
The team built all-DNA synthetic cells with a viscoelastic, crowded core and a permeable shell. Then, they assembled DNA nanotubes inside the core, forming artificial cytoskeletons anchored to the interior!
Their goal? To study self-assembly in crowded, viscoelastic environments, just like the cytoplasm. No diluted solutions here! Molecular crowding is well understood, but combined with viscoelasticity? Not so much.
DNA is perfect for this, because:
It’s programmable.
It self-assembles.
It can mimic key physical properties of cytoplasm.
Let’s see what they built!
Addressable DNA Droplets
The authors created SCs completely based on DNA.
These SCs are made from single-stranded DNA (ssDNA) polymers:
Adenine-rich strands (A-rich).
Thymine-rich strands (T-rich).
If you heat them together, the A-rich polymers condense into droplets, and the T-rich strands form a shell around them.
You end up with a core-shell structure:
Liquid, viscoelastic interior.
Permeable outer shell.
Super cool!
The droplets are completely addressable. Each strand carries a barcode that acts as a sequence-addressable handle:
k barcode in the T-rich shell.
m, p, s barcodes in enable orthogonal recruitment of specific components in the core.
This design clearly separates:
Physical compartment formation → core-shell droplet, based on phase-separation of A-rich and T-rich strands
Functional molecular recruitment → barcode-based, possible thanks to co-phase separation (two or more barcodes inside the core!)
Growing DNA Cytoskeletons
Once the droplets are built, the team introduces DNA tiles that assemble into DNA nanotubes that act as artificial cytoskeletons. The tiles bind to the m barcode in the core, anchoring them to the matrix. Then, they self-assemble into long, μm-scale tubes!
This way, the team can study self-assembly in a viscoelastic, crowded environment, similar to cells. But in this case, they can engineer everything:
Barcode concentration → defines the effective concentration.
Time → tubes can grow at different speeds.
Temperature → changes dynamics.
The most important factor? Barcode concentration.
A proper network only forms at intermediate concentrations. The matrix must be:
Dense enough to retain tiles
Not so concentrated that the tiles cannot grow into extended fibers.
Too low concentrations, and you get only short fibers, because there aren’t enough building blocks. Go too high, and there are no nanotubes because all the DNA tiles are bound to the matrix.
This is different from simple crowding. Here, the nanotubes are anchored and coupled to the viscoelastic interior, and cannot bundle together. And it suggests that growth proceeds by end-to-end joining of small fibers!
Stimuli-Controlled Cytoskeleton Formation
But the authors didn’t stop here.
They made their platform responsive:
Light-triggered assembly
The team designed a DNA tile with a photocleavable linker in a hairpin, blocking a sticky end. UV light breaks the linker, the hairpin opens, and the tile can self-assemble!DNAzyme-triggered assembly
Tile D is protected by a hairpin with a motif that is recognized and cleaved by a DNAzyme localized via the s barcode. Once the DNAzyme and tile D meet in the same SC, the hairpin is removed, triggering cytoskeleton formation.Strand-displacement control
A blocker strand inhibits Tile E. Add the activator strand: DNA strand displacement starts assembly! Add a deactivator: the cytoskeleton disappears. Fully reversible!Orthogonal cytoskeletons in the same compartment
Finally, the team built two orthogonal cytoskeletons in the same droplet, combining light-triggered and strand displacement. Depending on the stimulus combination, the fluorescent result can be:a single green cytoskeleton,
a single magenta cytoskeleton,
or both at once!
Functions: Stiffness and Cell Contacts
So, does this translate to functions?
The team used AFM to check mechanical stiffness. It turns out, SCs with DNA cytoskeletons are much stiffer than SCs without! And the elastic modulus increases to levels comparable to real cells.
This begs the question: can these SCs interact with normal cells?
To answer this, the team functionalized them with cRGD, an integrin-binding peptide, and incubated them with HeLa cells. Both SCs with and without DNA cytoskeletons deform after contacting cells.
The difference?
Without cytoskeleton → leak contents
With the cytoskeleton → stay intact
The cytoskeleton seems to improve mechanical durability!
Are DNA Cells Here?
Interesting idea!
It’s hard for me to think about cells as busy, mechanically constrained environments. Papers like these are a good reminder! This platform is interesting:
It’s programmable at different levels: Barcode tuning, selective enrichment, light activation, and more!
Real mechanical advantage: The DNA cytoskeleton reinforces the SCs; with the proper control, this can be switched on and off.
Modular and extensible design: They showed that it’s easy to add fluorescence and binders. What else could be added?
There is lots of work on soft microrobotics, with biomedical applications. Could these DNA droplets be a potential, programmable, and modular platform? It could be an interesting direction!
Plus, it’s amazing to see the collision of DNA nanotech and synthetic biology!
And if you want to learn more about viscoelasticity, the cytoskeleton, and how they interact, read the paper here!
If you made it this far, thank you! What do you think of synthetic cells? Do you think they have a place in biomedicine? Reply and let me know!
P.S: Know someone interested in DNA nanotech and SynBio? Share it with them!
What did you think of today's newsletter?
More Room:
DNA USB Sticks: The time for DNA-based USB sticks is almost here! DNA data storage is promising, but there are still problems to solve. In this paper, the authors develop a strategy for more accurate and unbiased DNA amplification in chip-scale DNA data storage. By optimizing primer–template hybridization, they enable reliable amplification of large DNA libraries, reducing sequencing costs and improving data retrieval. The system successfully stores and retrieves multimedia data with good accuracy even at very low sequencing depth, advancing scalable DNA-based data storage.
DNA Moving Machines: You know it, I’m a DNA nanotech fan. It’s just the coolest: you can create dynamic machines, motors, and switches that mimic biological systems! This review clarifies definitions between these devices and evaluates them using metrics like speed, force, and efficiency, comparing them to natural motors such as myosin and kinesin. It highlights design strategies like strand displacement and DNA origami, discusses applications in drug delivery and biosensing, and outlines key challenges in matching the performance of biological systems.
Assembling DNA at Room Temp: You know, one annoying thing with DNA nanotech, though? You need high-temperature ramps to assemble structures. Isothermal DNA assembly would simplify nanostructure formation, avoiding temperature cycling and enabling construction under physiological conditions. In this study, the authors demonstrate isothermal assembly of the PX DNA motif in different ions (Mg²⁺, Ca²⁺, Sr²⁺) at 20–37 °C, and show that assembly is thermodynamically favored at higher temperatures. They also incorporate thrombin-binding aptamers directly during assembly, highlighting the ability to attach functional biomolecules in a single-step, constant-temperature process.
